Usage¶
Start by importing healing_scattering_image.
import healing_scattering_image
Functions¶
-
healing_scattering_image.heal_scatter_image.heal_scatter_image(im, mask, xcenter, ycenter, r_min, r_max, angle_resolution, delta_width, bkgd_threshold, peaks_threshold, bins=30, lambda_threshold_1=1.0, lambda_threshold_2=1.0, bkgd_fit_bias=4.0, fold_bias=4.0, fittness_threshold=1.0, extreme_fitness=0.001, two_fold_apply=True, fittness_prior=True, down_sample=0.1, fitting_shift=True)[source]¶ Heal Scattering Image basing on Physical features. See paper: Liu, Jiliang, et al. “Healing X-ray scattering images.” IUCrJ 4.4 (2017): 455-465.
Due to qphi image will have very low azimuth resolution at low angle region, if there is pattern very close beam center, which pixel distance less than 40 pixel, we would like to up size sample first. Usually 500x500 or 1000x1000 size image could be well processedself. This is how default parameters or paratmeters in example determinedself.
Parameters: - im: 2D numpy.array
input scattering image with gaps and defects,
- mask: 2D numpy.array
the area want to mask in scattering image,, like beamstop, gaps,
- xcenter: pixel float
the beamcenter column,
- ycenter: pixel float
the beam center row,
- r_min: pixel float
the start of healing,
- r_max: pixel float
the end of healing,
- angle_resolution: float
angle resolution of qphi map,
- delta_width: float
delta q for standard deviation calultion of every q in qphi map
- bkgd_threshold: float
threshold to classify scattering by the histogram of local standard devation verse global deviation. larger threshold will be less senstive to
- peaks_threshold: float
threshold determines histogram of global standard deviation larger threshold means only sharp peaks are distinguishable,
- bins: float
bins for histogram of standard deviation
- lambda_threshold_1: float
lower threshold will allow more data point classified to patterns including anisotropic structrue information,
- lambda_threshold_2: float
lower threshold will allow more data point classified to patterns including anisotropic structrue information,
- bkgd_fit_bias: float
determine the bias to symmtry judgement of diffuse peaks,
- fold_bias: float
determine the bias to symmtry judgement of sharp peaks,
- fittness_threshold: float
increase error tolerance for diffuse reflections,
- extreme_fitness: float
tolerance of error of symmetr model,
- two_fold_apply: Boolean
automatical applied two fold symmetry, usualy true for SAXS,
- fittness_prior: Boolean
True will enable algorithm to choose larger symmetry fold with fitting residual less than extreme_fitness,
- down_sample: float
between (0,1),
- fitting_shift: Boolean
allow small shift when heal the symmetrical patterns.
Returns: - im: 2D numpy.array
healed scattering image,
- aniso_judge: Boolean
identify peaks with symmertical folds,
- sym_record: float
symmetrical fold,
- iso_judge_global: 1D numpy.array
global standard deviation,
- iso_local_vs_global: 1D nummpy.array
local vs global standard deviation,
- qphi_image_6: 2D numpy.array
healed qphi map,
- I: 1D numpy array
circular averaged intensity.
Example¶
In [1]: import numpy as np
In [2]: from skimage.draw import polygon
In [3]: import matplotlib.pyplot as plt
In [4]: from scipy.io import loadmat
In [5]: from scipy import ndimage
In [6]: from skimage.filters import median
In [7]: from skimage.transform import resize
In [8]: import healing_scattering_image
In [9]: from healing_scattering_image.heal_scatter_image import heal_scatter_image
In [10]: import os
In [11]: from pkg_resources import resource_filename
In [12]: path = resource_filename('healing_scattering_image',\
....: 'example_data/example_1.npz')
....:
In [13]: sample_name = path
In [14]: im = np.double(np.load(sample_name)['im'])
In [15]: im += 1.
In [16]: mask = np.load(sample_name)['mask']
In [17]: from skimage.transform import resize
#im = resize(im,(int(np.shape(im)[0]/2),int(np.shape(im)[1]/2)))
#mask = resize(mask,(int(np.shape(mask)[0]/2),int(np.shape(mask)[1]/2)))
In [18]: mask = mask.astype(bool)
In [19]: im[mask] = np.nan
In [20]: xcenter = (np.load(sample_name)['xcenter'])#/2
In [21]: ycenter = (np.load(sample_name)['ycenter'])#/2
In [22]: r_min = 10
In [23]: r_max = np.max(im.shape)#int(np.max(np.shape(im)))
In [24]: angle_resolution = 360
In [25]: delta_width = 2
In [26]: bkgd_threshold = 0.3
In [27]: peaks_threshold = 4
In [28]: healed_im, aniso_place, sym_record,\
In [28]: iso_judge_global , iso_local_vs_global,\
In [28]: qphi_image_6, I = heal_scatter_image(im,mask,
....: xcenter,
....: ycenter,
....: r_min,
....: r_max,
....: angle_resolution,
....: delta_width,
....: bkgd_threshold,
....: peaks_threshold,
....: bins = 30,
....: lambda_threshold_1 = 1.,
....: lambda_threshold_2 = 1.,
....: bkgd_fit_bias = 4.,
....: fold_bias = 6.,
....: fittness_threshold = 1.,
....: extreme_fitness = 1e-3,
....: two_fold_apply = True,
....: fittness_prior=True,
....: down_sample = 0.1,)
....:
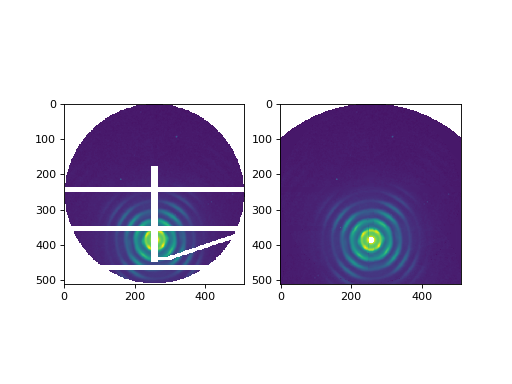
Plots¶